cellular computing and synthetic biology
Martyn Amos and Angel Goñi-Moreno survey the three "waves" of cellular computing approaches in synthetic biology over the last 3 decades.
Synthetic biology is founded on the idea that engineering principles can be used to modify and construct new (molecular) biological systems (sets of biological interactions that produce some coherent behavior). (This idea is new; molecular biology has long been more concerned with explaining the behavior of biological systems than it has been with creating new ones.)
Many see synthetic biology as an extension of genetic engineering, which examines how changes one or two genes affects the overall behavior of a cell or organism. By contrast, SynBio makes wholesale changes to biological systems, focusing on bottom-up construction of new systems. This practice has two goals:
Learn how biology works by recreating it synthetically.
Design biological systems that are more efficient than the ones that exist in nature. (Can we do better than evolution?)
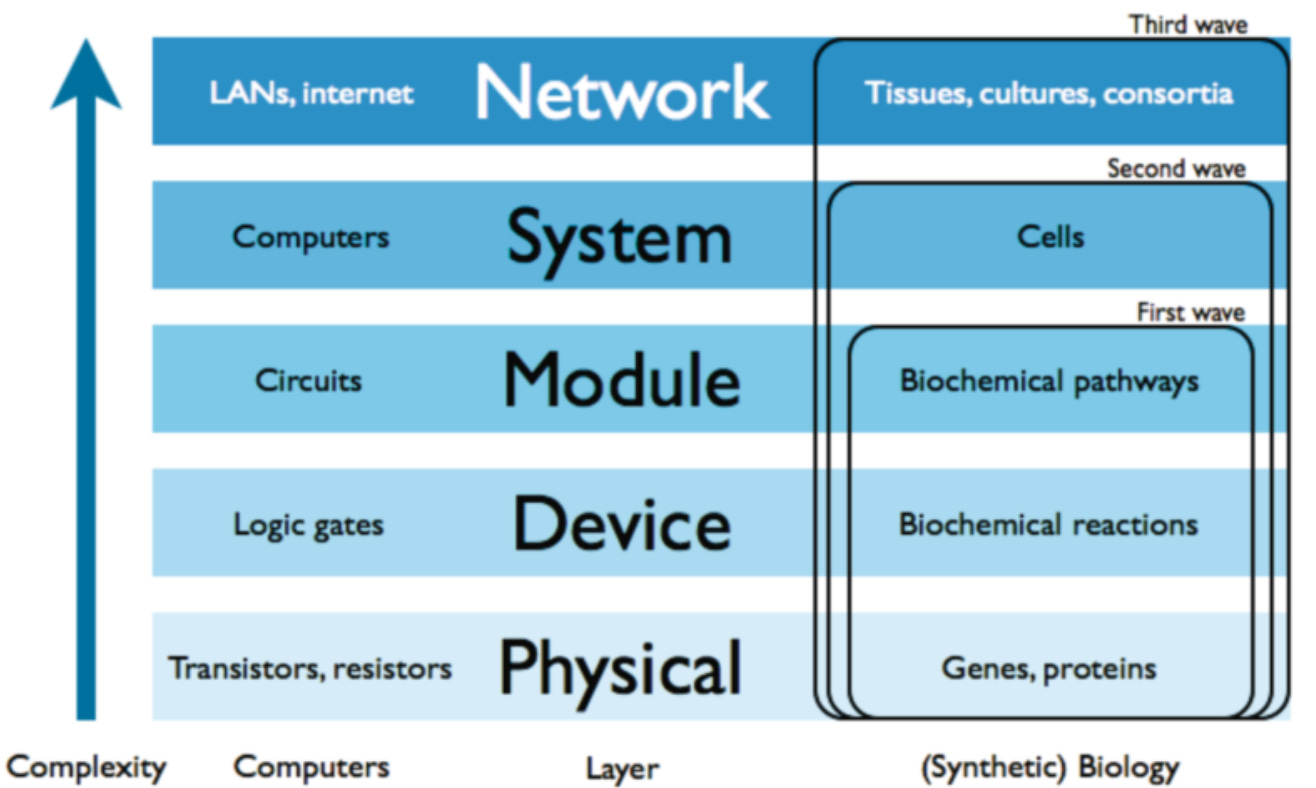
Implied in the goals of synthetic biology is the idea that biological systems store and process information in a way that is at least somewhat similar to computers. One of their compelling parallels is their hierarchical organization (see below).
(Fig. 7.1, Amos and Goñi-Moreno)
Working in such a hierarchically-organized way demands an ability for abstraction (capturing the essential features of a system at each level of complexity and hiding unnecessary details).
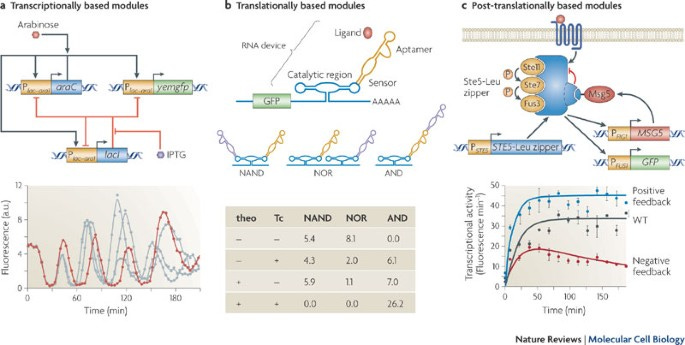
First-wave synthetic biology (around the early 2000s) worked at the modular level: different biochemical reactions were pieced together to form modules that could implement functions like oscillation and switching between two stable states.
(Fig. 1, https://www.nature.com/articles/nrm2698. From left to right: A module with genes that interact (by producing proteins that inhibit each other) to produce oscillations in a cell’s fluorescence level over time; Logic gates at the RNA level; Positive and negative feedback brought about by different ways of binding to a protein that induce changes in its function)
The second wave of SynBio brought modules together into systems: sets of modules which produce coherent behaviors. One idea that rose in popularity during this time was that of encapsulating biological modules in individual cells, and then working with these cells to produce system-level behaviors.
Cellular vs. silicon-based computing
Synthetic biology is filled with both enthusiasts and skeptics of cellular computing. “Every living cell within us is a hybrid analog-digital supercomputer that implements highly computationally-intensive nonlinear, stochastic, differential equations with 30,000 gene–protein state variables that interact via complex feedback loops,” writes Rahul Sarpeshkar at Dartmouth. (Cells are small, but mighty!)
On the other hand, the cellular environment is noisy and biological behavior is (at present) far less predictable than that of a classical computer. Does cellular computing hold promise?
Amos and Goñi-Moreno evaluate the potential success of cellular computing on the following criteria, among others (see Section 7.2 of their paper for the full list):
Speed: The authors conclude unequivocally that cells are outmatched when it comes to computation speed. They write, “cell-based computational devices will provide no competition for existing substrates on their “home turf”. Rather, such devices will find specific application niches, in which they might well outperform traditional silicon-based machines (for example, in the biosensing or medical device domains).” But there’s still hope: “In these types of situation, “speed” becomes a relatively meaningless metric.”
Resources:
Production: Designing and synthesizing nucleic acid constructs (i.e. engineered DNA) is a time- and resource-intensive process, but many expect these costs to go down by orders of magnitude over the next decade.
Computation: On this side, however, cells fare much better than traditional computers. Another note from Rahul Sarpeshkar: “The average 10 μm human cell performs these amazing computations with 0.34 nm self-aligned nanoscale DNA–protein devices, with 20 kT per molecular operation (1 ATP molecule hydrolysed), approximately 0.8 pW of power consumption (10 M ATP s−1) … Even at the end of Moore’s law, we will not match such performance by even a few orders of magnitude.”
Quality: Just how much we can do with cells remains to be seen. The success of SynBio relies on our ability to connect modules together to realize systems with increasingly complex, yet predictable and reliable, behavior.
Formalism: The field needs formal methods for the design and analysis of new biological systems. A formalized design process (instead of the case-by-case methods that are currently in use) will help the field along greatly.
Programmability: We need easy ways to dictate the high-level behavior of a biological system and have a program figure out the low-level implementation in the background (i.e. biological programming languages).
Summary + the third wave of SynBio
Though it still lacks a formal definition, synthetic biology is generally guided by these principles:
Designing biological systems to meet predefined user specifications.
Modularity
Abstraction
By treating cells as microscopic computers, we can translate knowledge and formalisms from computer science into synthetic biology.
The authors anticipate that third-wave synthetic biology will see a shift from systems to network-level engineering: “We distinguish ‘systems’ from ‘networks’ by measuring their capacity for information transmission; within systems we generally observe relatively low rates of transmission, whereas networks have much higher ‘bandwidth’ capabilities,” they write. They also predict a shift from engineering at the level of single cells to multi-cellular approaches.
For all of this to happen, the field will need close collaboration between biologists, mathematicians, computer scientists, and engineers. In under three decades, synthetic biology has made huge advances towards its goals of rationally engineering biology, and experts in fields from science to economics predict equally good progress to come.
Find the article here:
Amos, M., & Goñi-Moreno, A. (2018). Cellular Computing and Synthetic Biology. In S. Stepney, S. Rasmussen, & M. Amos (Eds.), Computational Matter (pp. 93–110). Springer International Publishing. https://doi.org/10.1007/978-3-319-65826-1_7