parallel computing with DNA
A DNA-based solution to the directed Hamiltonian parallel path problem.
Leonard Adleman is widely recognized as the father of ‘DNA computing,’ an approach that uses DNA molecules and biochemical reactions to store and process information. (He is better known for RSA (Rivest-Shamir-Adleman) encryption, a system which he developed in 1977 that continues to be widely used for secure data transmission.)
The problem
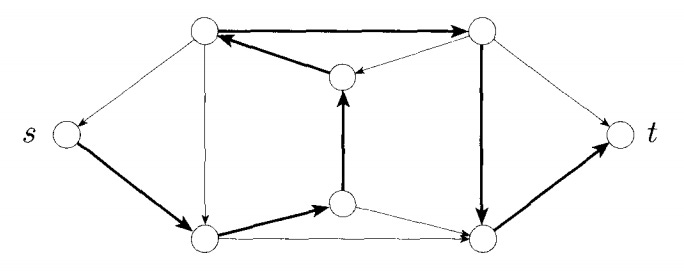
Given a graph with directed edges and selected “start” and “end” nodes, find a “Hamiltonian path:” a path that goes from “start” to “end,” visiting each node in the graph exactly once.
A directed Hamiltonian path (s = start, t = end). (Ian Finlayson)
This problem has been shown to be NP-complete: while solutions to it can be verified easily, there is no known (deterministic) way to quickly find a solution.
Adleman’s solution
Leonard Adleman proposes a nondeterministic algorithm to solve the directed Hamiltonian path problem, then shows how it can be implemented using DNA.
Algorithm:
Randomly generate paths through the graph (can start and end at any node and be of any length).
Take out the paths that do not start and end at the designated start and end nodes.
Take out the paths that do not visit exactly the same number of nodes as there are in the graph (remember that a directed Hamiltonian path visits every node in the graph exactly once).
Take out the paths that do not visit every node in the graph.
At this point, if any paths remain, they are directed Hamiltonian paths. If not, the graph does not have any directed Hamiltonian paths.
Implementing in DNA:
Step 1: Forming random paths.
Represent each node N in the graph with a random 20 bp sequence of DNA (sort of like an ID). (Node A gets DNAA.) Take the complement of all of the node DNA sequences (/DNAN).
Represent each directed edge EI—>J in the graph with a 20 bp DNA sequence: the last 10 bp of DNAA followed by the first 10 bp of DNAB. (Edge EA—>B gets DNAA—>B).
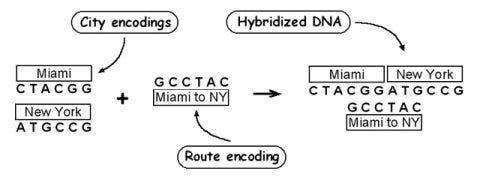
To make random paths, mix the complement node DNA sequences, /DNAN, with the edge sequences, EI—>J, in a ligation reaction (a reaction that joins DNA fragments). The node and edge sequences bind together (remember complementary binding!), with each edge sequence serving as a link between two nodes, and are ligated by the enzymes in the reaction solution. At the end of this process, we have random paths of varying lengths.
Generating a path from ‘Miami’ to ‘New York’ using complementary base pair binding. (Ars Technica)
Step 2: Taking out paths that do not start and end at the designated nodes.
Paths that start and end at specific nodes can be selected using PCR, a DNA amplification reaction. PCR works by mixing short DNA sequences called primers, into a sample containing many different DNA fragments. The primers are designed to bind only to the sequence we are interested in amplifying. After primer binding, enzymes copy the DNA sequence following the primer, creating a (complementary) copy of the target DNA. By repeating this process 20-40 times, the DNA sequence of interest is amplified (i.e. there are many copies of it relative to all other DNA fragments in the sample).
PCR amplification (Khan Academy).
Here, we choose our primers to be the 20bp sequences representing the start and end nodes. All of the paths that are amplified with PCR then must start and end with the designated nodes.
Step 3: Taking out paths that are too long or short.
Gel electrophoresis1 is used to separate DNA fragments by length. If we know how many nodes N are in the graph, we can estimate the length of DNA fragments representing a path that is N nodes long (20*N). The band in the gel corresponding to fragments that are 20N bp long is removed and the DNA within is purified.
Step 4: Taking out paths that do not visit all nodes.
One by one, the sample is mixed with node DNA fragments that are bound to small magnetic beads. When DNA1 (with magnetic beads) is added, it binds only to paths that contain the sequence for node 1. Using a magnetic field, these paths can be filtered out from the sample. This process is repeated for all nodes in the graph.
The paths that now remain are the ones that:
start and end at the desired locations.
are exactly N nodes long.
visit all nodes.
… so they must be directed Hamiltonian paths. (If nothing is left, there are no directed Hamiltonian paths).
Benefits and drawbacks
For the most part, this workflow uses basic molecular biology techniques (it’s nothing fancy!). It is massively parallel: we can check almost all paths through the graph at once. It is also highly energy efficient - 1 J is enough for about 2*1019 ‘operations’ (ligation of two DNA molecules, or sequence amplification). (Supercomputers (at the time this was written) could execute at most 109 operations per joule.) Information density is also high, at about 1 bit per cubic nanometer. By contrast, videotapes hold about 1 bit per 1012 cubic nanometers.
However, lab work is time- and labor-intensive, and computing solutions for a 6-node graph in this paper took about 7 days. Synthesizing sequences beforehand to represent the nodes and edges is also a time-consuming process. We should additionally consider error rates in each of these steps: for example, imperfect or incorrect binding between node and edge DNAs in Step 1 could result in ‘pseudopaths,’ paths that do not actually occur, which could produce misleading results.
Adleman concludes that molecular computation might become useful if we can overcome the amount of manual lab work required to implement it. While DNA computing will not replace silicon for operations like arithmetic or executing deterministic algorithms, it might compete with or even surpass electronic computation (an older post about this here) on particular problems. DNA computing’s biggest selling point is parallelism, and this feature can be exploited by finding appropriate problems that parallel, nondeterministic algorithms can efficiently solve.
Find the paper here:
Adleman, L. M. (1994). Molecular Computation of Solutions to Combinatorial Problems. Science, 266(5187), 1021–1024. https://doi.org/10.1126/science.7973651
Gel electrophoresis separates DNA molecules based on their size and electrical charge by forcing them through a gel matrix using an electric current.